3D genome organization and GRiNCH with Da-Inn Erika Lee (#61)
June 23, 2021
In this episode, Jacob Schreiber interviews Da-Inn Erika Lee about data and computational methods for making sense of 3D genome structure. They begin their discussion by talking about 3D genome structure at a high level and the challenges in working with such data. Then, they discuss a method recently developed by Erika, named GRiNCH, that mines this data to identify spans of the genome that cluster together in 3D space and potentially help control gene regulation.
Links:
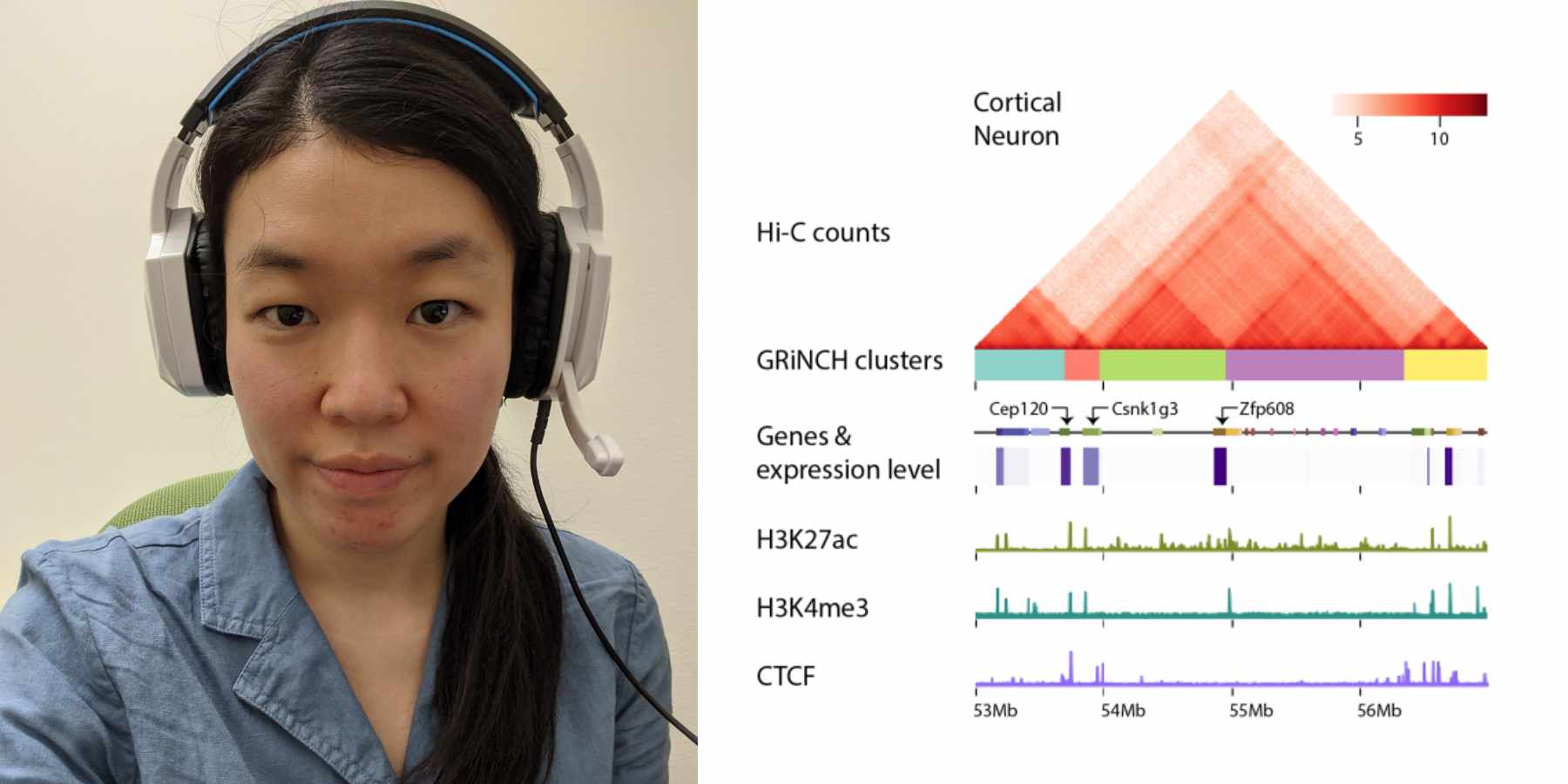
- GRiNCH: simultaneous smoothing and detection of topological units of genome organization from sparse chromatin contact count matrices with matrix factorization (Da-Inn Lee and Sushmita Roy)
- GRiNCH Project Page
- In silico prediction of high-resolution Hi-C interaction matrices (Shilu Zhang, Deborah Chasman, Sara Knaack, and Sushmita Roy)
Music: Eric Skiff — Come and Find Me (modified, licensed under CC BY 4.0).
Subscribe to the bioinformatics chat on Apple Podcasts, Pocket Casts, Spotify, or any other podcasting app via the RSS feed link. You can also follow the podcast on Mastodon and Twitter and support it on Patreon.
