Seeding methods for read alignment with Markus Schmidt (#54)
December 16, 2020
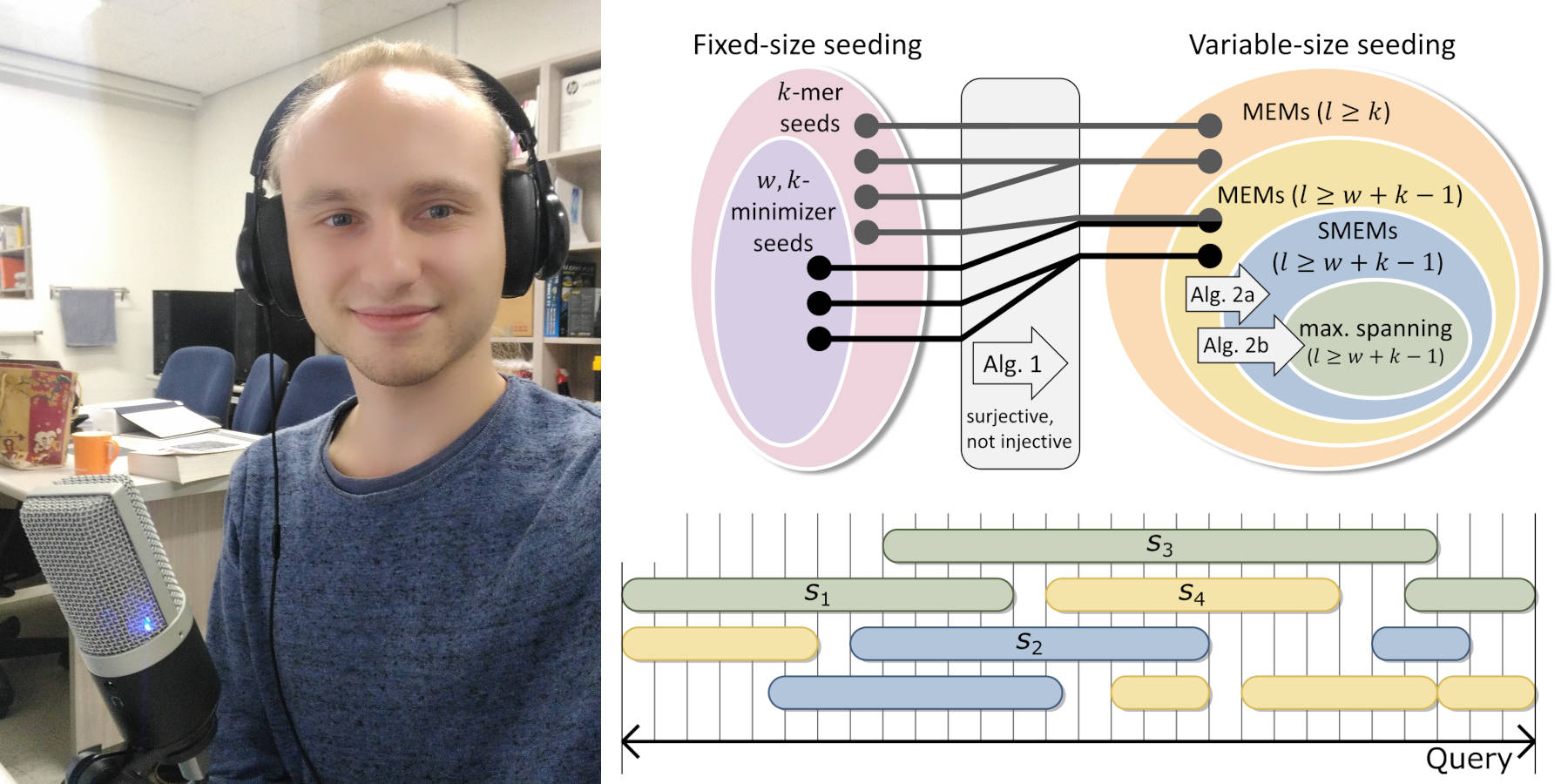
In this episode, Markus Schmidt explains how seeding in read alignment works. We define and compare k-mers, minimizers, MEMs, SMEMs, and maximal spanning seeds. Markus also presents his recent work on computing variable-sized seeds (MEMs, SMEMs, and maximal spanning seeds) from fixed-sized seeds (k-mers and minimizers) and his Modular Aligner.
Links:
- A performant bridge between fixed-size and variable-size seeding (Arne Kutzner, Pok-Son Kim, Markus Schmidt)
- MA the Modular Aligner
- Calibrating Seed-Based Heuristics to Map Short Reads With Sesame (Guillaume J. Filion, Ruggero Cortini, Eduard Zorita) — another interesting recent work on seeding methods (though we didn’t get to discuss it in this episode)
Music: Eric Skiff — Come and Find Me (modified, licensed under CC BY 4.0).
Subscribe to the bioinformatics chat on Apple Podcasts, Pocket Casts, Spotify, or any other podcasting app via the RSS feed link. You can also follow the podcast on Mastodon and Twitter and support it on Patreon.
